Genome Assembly (de novo)
Genome Assembly (reference-based)
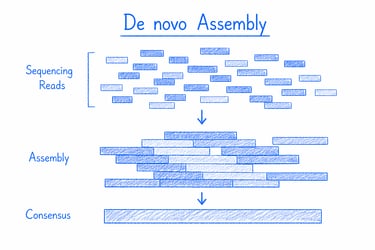
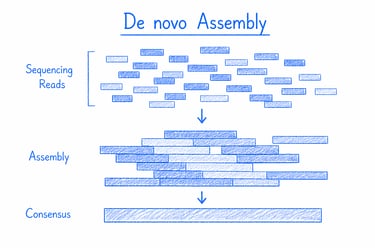
De novo genome assembly reconstructs a complete genome directly from raw sequencing data, enabling the discovery of novel organisms and unique genetic variations. It’s ideal for unlocking new species' potential and uncovering hidden biological insights for research and biotech innovation.
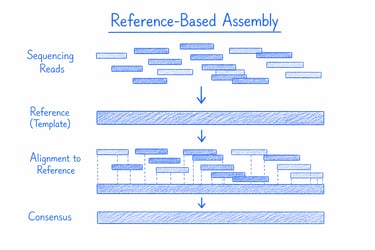
Reference-based genome assembly reconstructs a genome by aligning sequencing reads to an existing reference, delivering fast and accurate results for well-characterised organisms. It’s ideal for variant detection, comparative genomics, and large-scale studies where precision and efficiency are key.
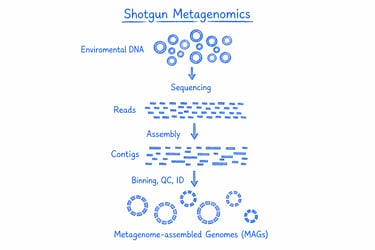
Shotgun metagenomics sequences all genetic material in a sample, giving you a comprehensive view of the entire microbial community in a single run. It’s ideal for exploring biodiversity, uncovering functional potential, and gaining deep insights into complex ecosystems for research and biotech applications.
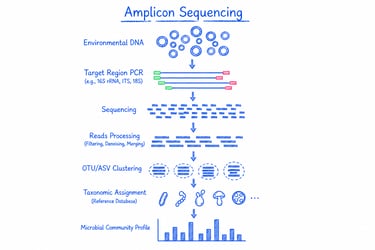
Amplicon sequencing targets and sequences specific genetic markers (such as 16S rRNA, ITS, or 18S) to profile microbial communities with high sensitivity and cost efficiency. It’s ideal for microbial diversity studies, taxonomic identification, community comparison, and routine microbiome analysis.
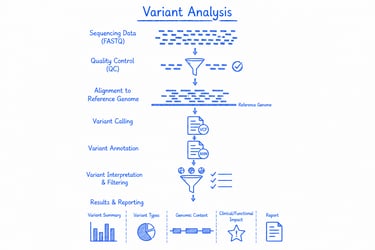
Variant analysis detects genetic variations (such as SNPs, indels, and copy number variations) by comparing sequencing data to a reference genome. It’s ideal for genotype profiling, disease research, and biomarker discovery.