Whole Transcriptomic Analysis
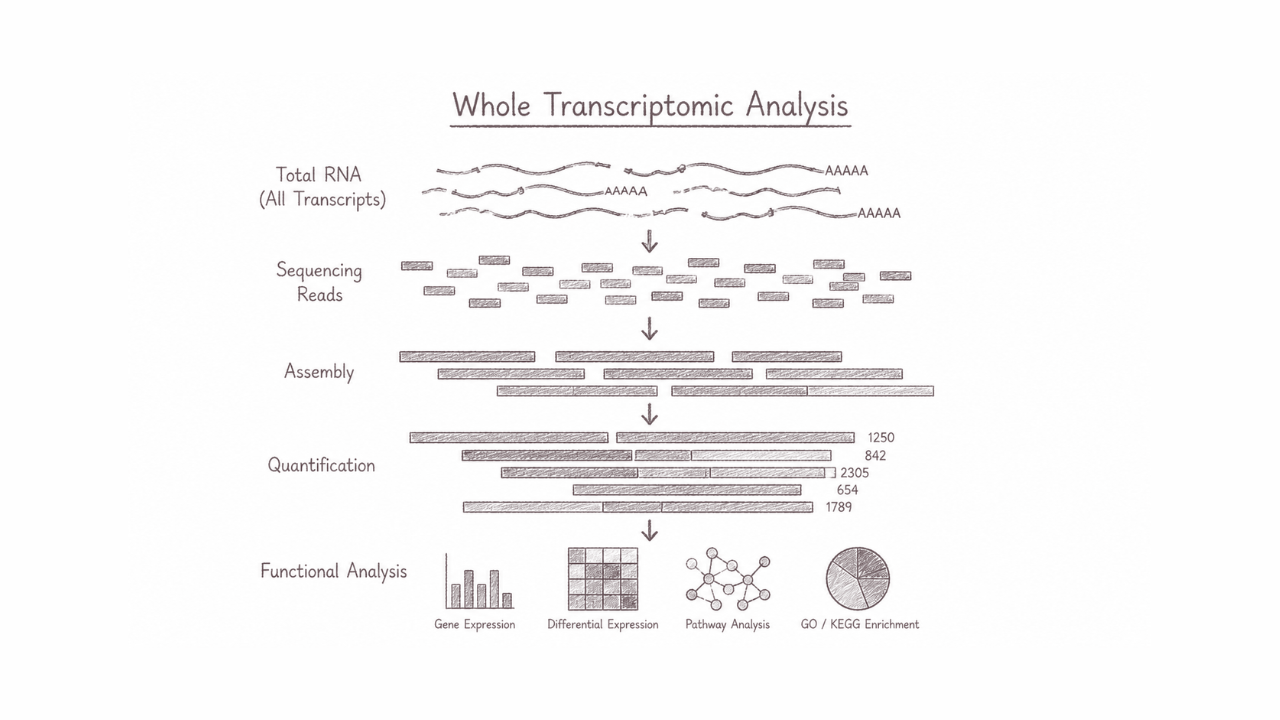
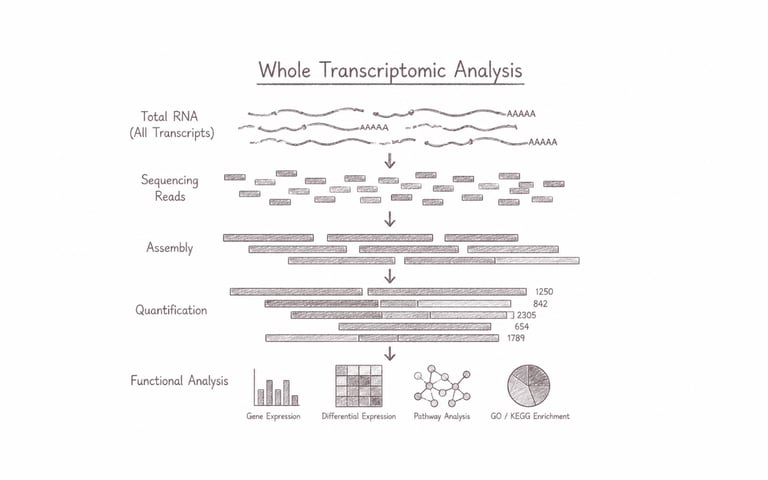
Whole transcriptomic analysis comprehensively profiles the complete set of RNA transcripts expressed within a cell, tissue, or organism under specific conditions. This approach captures both coding and non-coding RNA dynamics, enabling researchers to uncover gene expression patterns, regulatory networks, alternative splicing events, and functional biological responses at a system-wide scale. It is particularly powerful for understanding cellular mechanisms, disease pathways, developmental biology, and discovering molecular biomarkers that drive advances in precision medicine and biotechnology.
Base Pricing & Turnaround Time
Start from: Rp. 1.200.000 ($ 70) per sample
Start from: 7 days
Additional charges may apply if the FASTQ file size is unusually large
Default Deliverables
Quality-Controlled Sequence Files
FASTA/FASTQ files containing filtered, trimmed, and quality-controlled RNA sequencing reads generated from raw transcriptomic sequencing data before downstream expression analysis.
Aligned Transcript Read Files
BAM/CRAM files containing RNA sequencing reads aligned to the selected reference genome or transcriptome, including indexed alignment files for transcript visualisation and downstream analysis.
Transcript Assembly & Quantification Files
Transcript assembly and abundance estimation files containing reconstructed transcripts, transcript isoforms, and quantified gene/transcript expression levels generated from RNA sequencing data.
Differential Gene Expression Report
Comprehensive analysis of significantly upregulated and downregulated genes across experimental conditions, including statistical significance, fold-change values, and expression profiling results.
Functional Enrichment Analysis
Biological interpretation of expressed and differentially expressed genes, including Gene Ontology (GO), KEGG pathway enrichment, and functional category analysis.
Add-ons
Alternative Splicing Analysis
Novel Transcript Discovery
Fusion Gene Detection
Non-coding RNA Analysis
Pathway Activity Analysis
Co-expression Network Analysis
Time-Series Expression Analysis
Host–Pathogen Transcriptomic Interaction Analysis
Single-Sample Expression Profiling
