Metatranscriptomic Analysis
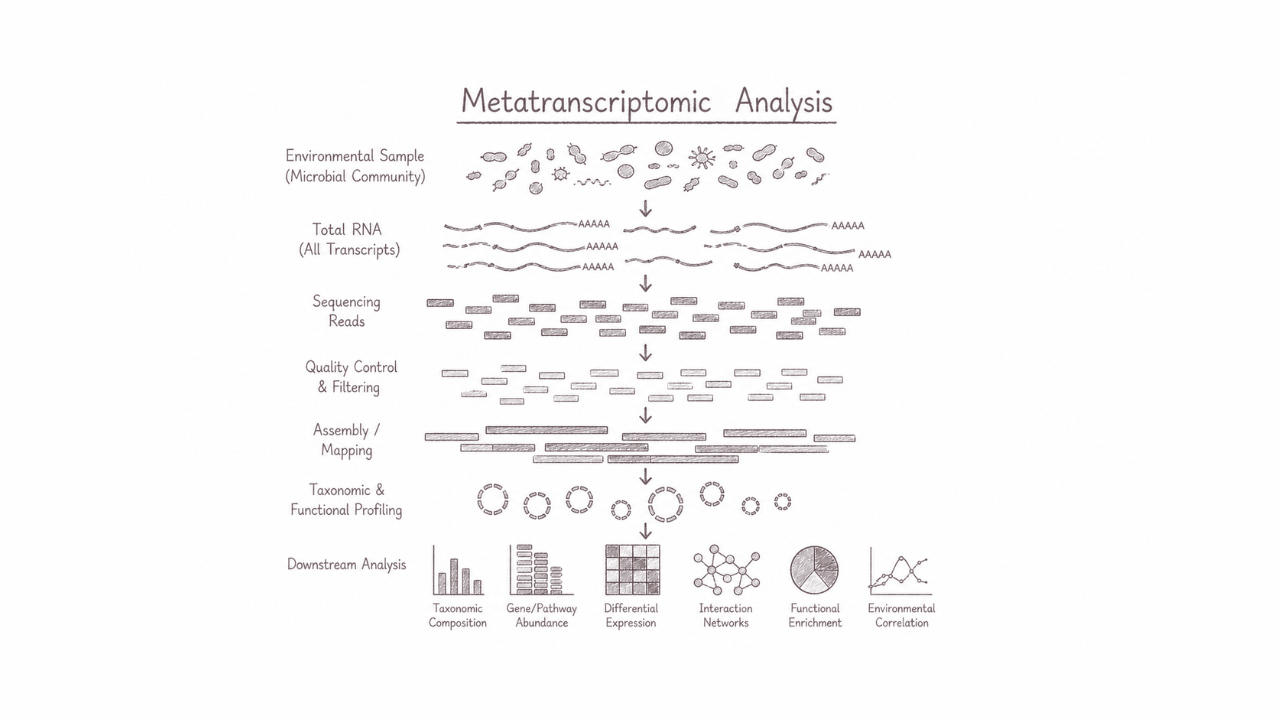
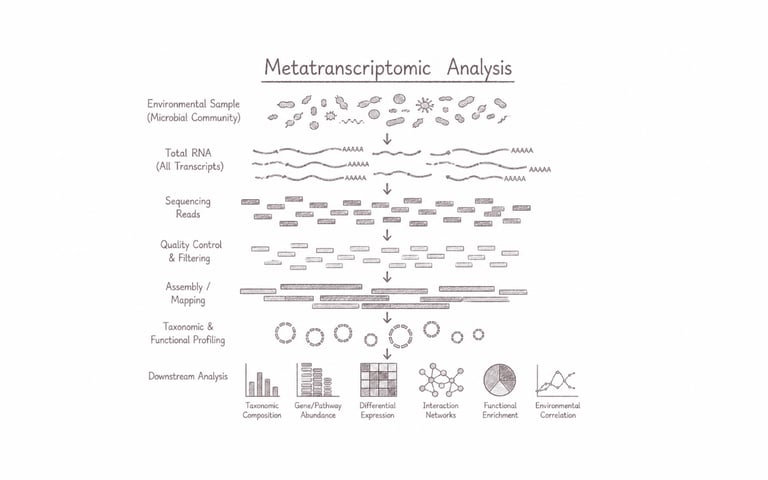
Metatranscriptomic analysis explores the complete collection of actively expressed RNA transcripts within complex microbial communities directly from environmental or host-associated samples. This approach reveals real-time functional activity, metabolic interactions, and dynamic responses of microorganisms without the need for cultivation. It is particularly powerful for understanding microbiome functionality, host–microbe interactions, ecosystem processes, and discovering active biological pathways that drive environmental adaptation, human health, and biotechnological innovation.
Base Pricing & Turnaround Time
Start from: Rp. 1.200.000 ($ 70) per sample
Start from: 7 days
Additional charges may apply if the FASTQ file size is unusually large
Default Deliverables
Quality-Controlled Sequence Files
FASTA/FASTQ files containing filtered, trimmed, and quality-controlled metatranscriptomic sequencing reads generated from raw environmental or host-associated RNA sequencing data before downstream functional analysis.
Host & Contaminant Removal Files
Processed sequencing files generated after removal of host-derived, ribosomal RNA (rRNA), and potential contaminant sequences to enrich actively expressed microbial transcripts.
Functional Gene Expression Analysis
Quantitative analysis of actively expressed microbial genes and metabolic functions, including pathway activity, enzyme classification, and community-level functional profiling.
Differential Functional Activity Report
Comparative analysis of microbial transcriptional activity across samples or conditions, including differentially expressed functional pathways, genes, and metabolic responses.
Add-ons
Novel Transcript Discovery
Fusion Gene Detection
Non-coding RNA Analysis
Pathway Activity Analysis
Co-expression Network Analysis
Time-Series Expression Analysis
Host–Pathogen Transcriptomic Interaction Analysis
Time-series Transcriptomic Analysis
