Spatial Transcriptomics Analysis
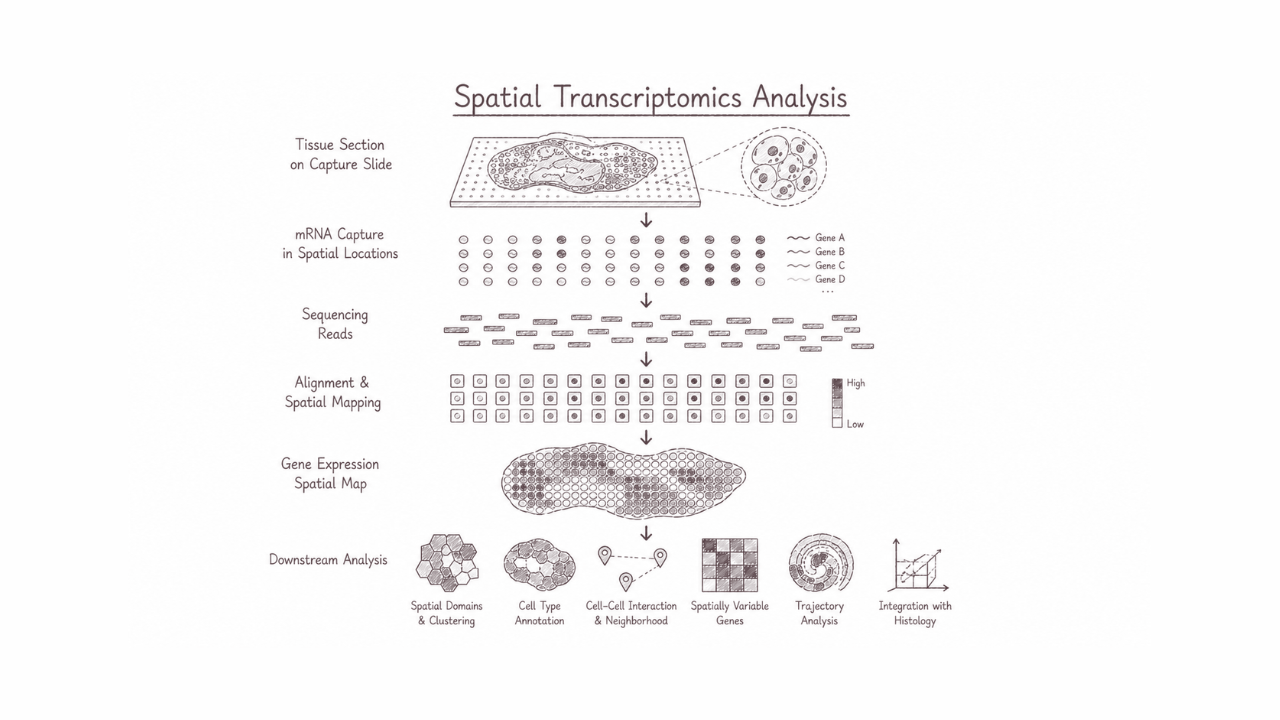
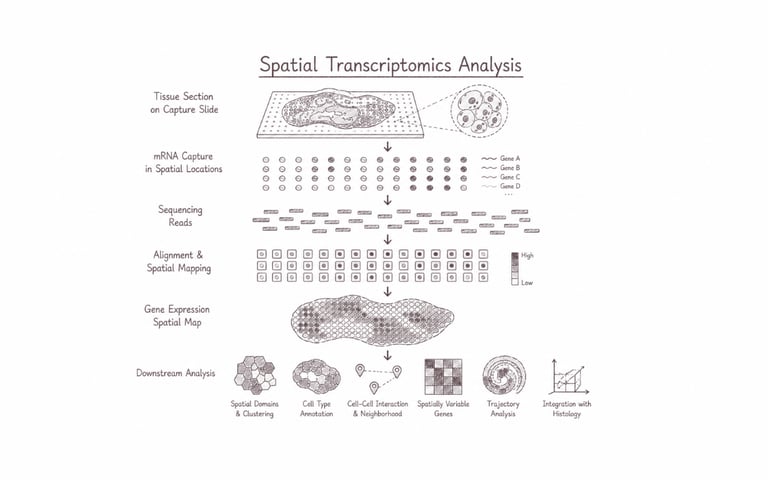
Spatial transcriptomics analysis maps gene expression directly within intact tissue architecture, preserving the spatial context of cellular activity and interactions. This approach enables researchers to visualise where specific transcripts are expressed, uncover cellular heterogeneity, tissue organisation, and localised molecular responses that conventional transcriptomics may overlook. It is particularly powerful for studying tumour microenvironments, developmental processes, neuroscience, and complex tissue biology, advancing deeper insights into how spatial cellular dynamics shape health and disease.
Base Pricing & Turnaround Time
Start from: Rp. 3.000.000 ($ 180) per sample
Start from: 7 days
Additional charges may apply if the FASTQ file size is unusually large
Default Deliverables
Spatially Resolved Gene Expression Matrix
Comprehensive spatial transcriptomic expression matrices containing transcript abundance data linked to precise tissue coordinates or capture spots across the analysed sample.
Spatial Barcode & Coordinate Mapping Files
Processed spatial indexing files containing barcode assignments, positional coordinates, and spot-to-tissue mapping information required for downstream spatial reconstruction and visualisation.
Tissue Image Processing & Registration
Aligned and processed histological or fluorescence tissue images integrated with spatial transcriptomic data for accurate molecular-to-tissue correspondence.
Spatial Clustering & Tissue Region Identification
Identification of transcriptionally distinct spatial domains, cellular neighbourhoods, or tissue regions based on localised gene expression patterns across the sample.
Spatially Variable Gene Analysis
Detection of genes exhibiting significant spatial expression variability, highlighting location-specific molecular activity and tissue-associated transcriptional programs.
Cellular Interaction & Spatial Network Analysis
Inference of spatial cellular communication patterns, neighbouring cell relationships, and local microenvironment interactions within the tissue architecture.
Add-ons
Cell Type Deconvolution
Trajectory & Spatial Pseudotime Analysis
Spatial Multi-Omics Integration
